 Research Areas > Beach Water Quality > Side by Side Beta-Testing of Rapid Methods
Research Areas > Beach Water Quality > Side by Side Beta-Testing of Rapid Methods
Project: Side-by-Side Beta Testing of Rapid Methods
Background and Objectives
Public health officials routinely monitor fecal indicator bacteria levels to assess beach water quality, but current laboratory processing methods generally require 24 hours for completion. In the meantime, swimmers can be exposed to poor water quality. Several researchers have developed new methods that produce results within two hours, presenting a need for independent testing to evaluate the performance of the new methods. SCCWRP previously conducted two evaluation studies in which methods were implemented by the developers. This project entails a third evaluation study in which practitioners from local laboratories were asked to apply two of the promising methods.
Status
This study was conducted in 2006.
Methods
The study involved simultaneous processing of samples using both rapid methods and existing methods to enumerate fecal indicator bacteria. The rapid methods tested were Quantitative Polymerase Chain Reaction (QPCR) and Transcription-Mediated Amplification (TMA). The existing methods included EPA Method 1600 (mEI agar) and the IDEXX defined substrate method. All samples were processed in duplicate using both types of methods.
QPCR was performed for both enterococci and Escherichia coli by the Orange County Sanitation District. TMA was performed for enterococci by the Orange County Public Health Agency. One hundred sixty-three samples were processed, 138 of which were ambient samples collected from 41 locations. The remaining samples consisted of clean seawater spiked with primary sewage influent or secondary effluent. A small set of additional samples was initially inoculated with secondary sewage effluent and then treated with chlorine that was subsequently neutralized. These were analyzed at several time points over the course of several hours in order to assess how well the rapid methods could enumerate enterococci in chlorine-disinfected wastewater.
Each method’s performance was evaluated with respect to the State’s water quality standards and for variability among replicates. Lastly, the evaluations were integrated into an overall assessment to determine if management decisions based on rapid method results would have been the same as those based on traditional methods.
Findings
• Technology transfer to working laboratories was successful, as laboratory personnel with minimal previous experience in performing molecular methods were able to master the rapid techniques in a relatively short period of time.
• Both rapid methods were more than 85% accurate with respect to the state standard for enterococci.
• Both rapid methods exhibited average coefficients of variation comparable to those of existing methods.
• The rapid and existing methods produced results for enterococci that would have led a public health officer to make the same beach management decision approximately 70% of the time.
• Rapid methods tended to underestimate bacterial concentrations as compared to traditional methods, primarily in stormdrain samples.
• In general, underestimation appeared to be due to inhibition of DNA amplification (caused by inhibitory substances in the water sample). Method developers are committed to future inclusion of new internal controls in the assays that will detect amplification inhibition and provide a correction factor if it occurs.
• Both rapid methods overestimated bacterial concentrations in the chlorinated effluent samples.
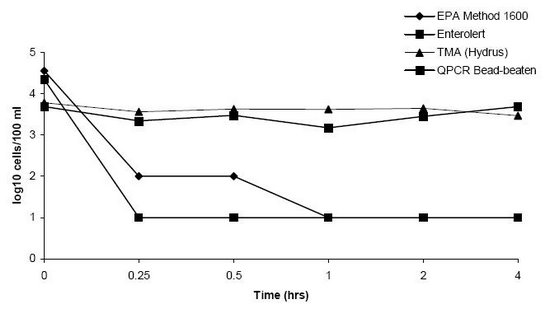
Effect of chlorination of levels of enterococci in sterile water spiked with secondary sewage effluent as measured by EPA-approved and rapid methods.
Partners
This study was conducted in collaboration with the Orange County Public Health Agency, Orange County Sanitation District, and the California Beach Water Quality Work Group.
This page was last updated on: 7/2/2014